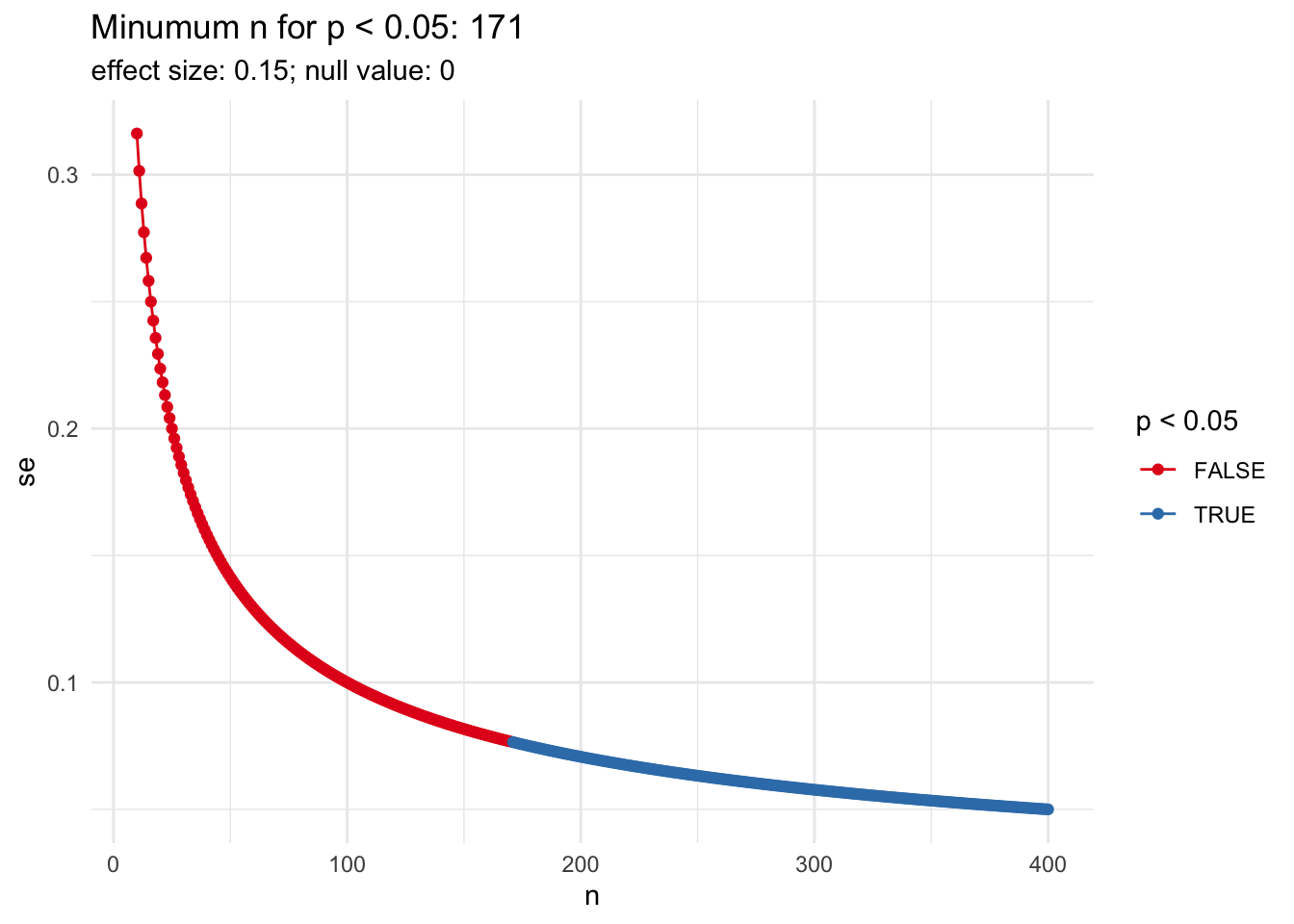
#' Calculate p-value from a chi-squared test with varying sample sizes
#'
#' This algorithm will start with an initial sample size (`n_start`) and perform a chi-squared test
#' with a vector of counts equal to `n * probs`. This will repeat increasing the sample size by
#' `n_step` until the p-value from the chi-squared test is less than `p_stop`.
#'
#' @param vector of cell probabilities. The sum of the values must equal 1.
#' @param sig_level significance level.
#' @param p_stop the p-value to stop estimating chi-squared tests.
#' @param max_n maximum n to attempt if `p_value` is never less than `p_stop`.
#' @param min_cell_size minimum size per cell to perform the chi-square test.
#' @param n_start the starting sample size.
#' @param n_step the increment for each iteration.
#' @return a data.frame with three columns: n (sample size), p_value, and sig (TRUE if
#' p_value < sig_level).
#' @importFrom DescTools power.chisq.test CramerV
chi_squared_power <- function(
probs,
sig_level = 0.05,
p_stop = 0.01,
power = 0.80,
power_stop = 0.90,
max_n = 100000,
min_cell_size = 10,
n_start = 10,
n_step = 10
) {
if(sum(probs) != 1) { # Make sure the sum is equal to 1
stop('The sum of the probabilities must equal 1.')
} else if(length(unique(probs)) == 1) {
stop('All the probabilities are equal.')
}
n <- n_start
p_values <- numeric()
power_values <- numeric()
df <- ifelse(is.vector(probs),
length(probs) - 1,
min(dim(probs)) - 1) # Degrees of freedom
repeat {
x <- (probs * n) |> round()
if(all(x > min_cell_size)) {
cs <- chisq.test(x, rescale.p = TRUE, simulate.p.value = FALSE)
p_values[n / n_step] <- cs$p.value
pow <- DescTools::power.chisq.test(
n = n,
w = DescTools::CramerV(as.table(x)),
df = df,
sig.level = sig_level
)
power_values[n / n_step] <- pow$power
if((cs$p.value < p_stop & pow$power > power_stop) | n > max_n) {
break;
}
} else {
p_values[n / n_step] <- NA
power_values[n / n_step] <- NA
}
n <- n + n_step
}
result <- data.frame(n = seq(10, length(p_values) * n_step, n_step),
p_value = p_values,
sig = p_values < sig_level,
power = power_values)
class(result) <- c('chisqpower', 'data.frame')
attr(result, 'probs') <- probs
attr(result, 'sig_level') <- sig_level
attr(result, 'p_stop') <- p_stop
attr(result, 'power') <- power
attr(result, 'power_stop') <- power_stop
attr(result, 'max_n') <- max_n
attr(result, 'n_step') <- n_step
return(result)
}
#' Plot the results of chi-squared power estimation
#'
#' @param x result of [chi_squared_power()].
#' @param plot_power whether to plot the power curve.
#' @param plot_p whether to plot p-values.
#' @param digits number of digits to round to.
#' @param segement_color color of the lines marking where power and p values exceed threshold.
#' @param sgement_linetype linetype of the lines marking where power and p values exceed threshold.
#' @param p_linetype linetype for the p-values.
#' @param power_linetype linetype for the power values.
#' @param title plot title. If missing a title will be automatically generated.
#' @parma ... currently not used.
#' @return a ggplot2 expression.
plot.chisqpower <- function(
x,
plot_power = TRUE,
plot_p = TRUE,
digits = 4,
segment_color = 'grey60',
segment_linetype = 1,
p_linetype = 1,
power_linetype = 2,
title,
...
) {
pow <- attr(x, 'power')
p <- ggplot(x[!is.na(x$p_value),], aes(x = n, y = p_value))
if(plot_power) {
if(any(x$power > pow, na.rm = TRUE)) {
min_n_power <- min(x[x$power > pow,]$n, na.rm = TRUE)
p <- p +
geom_segment(
x = 0,
xend = min_n_power,
y = pow,
yend = pow,
color = segment_color,
linetype = segment_linetype) +
ggplot2::annotate(
geom = 'text',
x = 0,
y = pow,
label = paste0('Power = ', pow),
vjust = -1,
hjust = 0) +
geom_segment(
x = min_n_power,
xend = min_n_power,
y = pow,
yend = 0,
color = segment_color,
linetype = segment_linetype) +
ggplot2::annotate(
geom = 'text',
x = min_n_power, y = 0,
label = paste0('n = ', prettyNum(min_n_power, big.mark = ',')),
vjust = 1,
hjust = -0.1)
}
p <- p +
geom_path(
aes(y = power),
color = '#7570b3',
linetype = power_linetype)
}
if(plot_p) {
if(any(x$sig, na.rm = TRUE)) {
p <- p +
geom_segment(
x = 0,
xend = min(x[x$sig,]$n, na.rm = TRUE),
y = attr(x, 'sig_level'),
yend = attr(x, 'sig_level'),
color = segment_color,
linetype = segment_linetype) +
ggplot2::annotate(
geom = 'text',
x = 0,
y = attr(x, 'sig_level'),
label = paste0('p = ', attr(x, 'sig_level')),
vjust = -1,
hjust = 0) +
geom_segment(
x = min(x[x$sig,]$n, na.rm = TRUE),
xend = min(x[x$sig,]$n, na.rm = TRUE),
y = attr(x, 'sig_level'),
yend = 0,
color = segment_color,
linetype = segment_linetype) +
ggplot2::annotate(
geom = 'text',
x = min(x[x$sig,]$n, na.rm = TRUE),
y = 0,
label = paste0('n = ', prettyNum(min(x[x$sig,]$n, na.rm = TRUE), big.mark = ',')),
vjust = 1,
hjust = -0.1)
}
p <- p +
geom_path(
alpha = 0.7,
linetype = p_linetype)
# geom_point(aes(color = sig), size = 1) +
# scale_color_brewer(paste0('p < ', attr(x, 'sig_level')), type = 'qual', palette = 6)
}
if(missing(title)) {
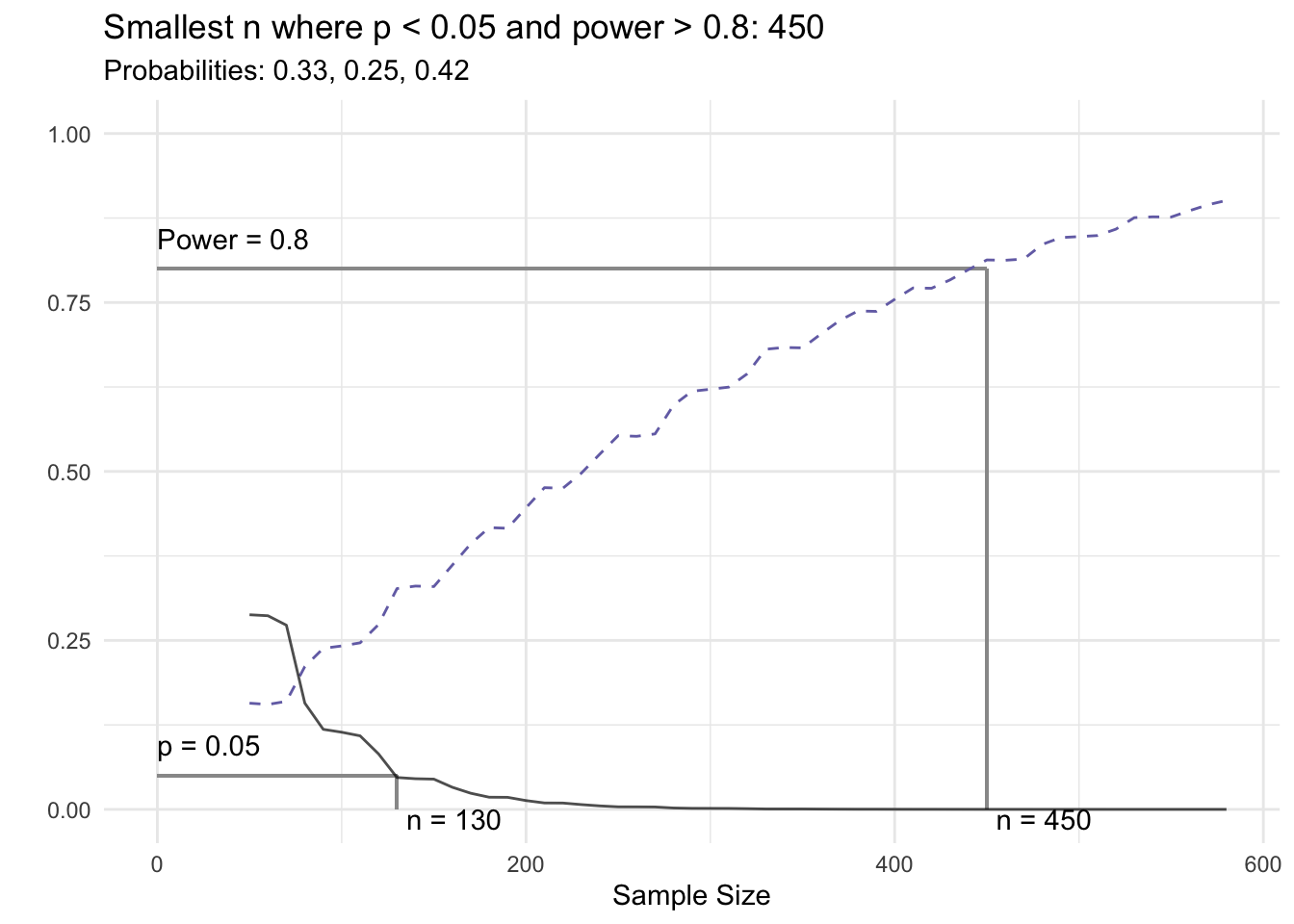
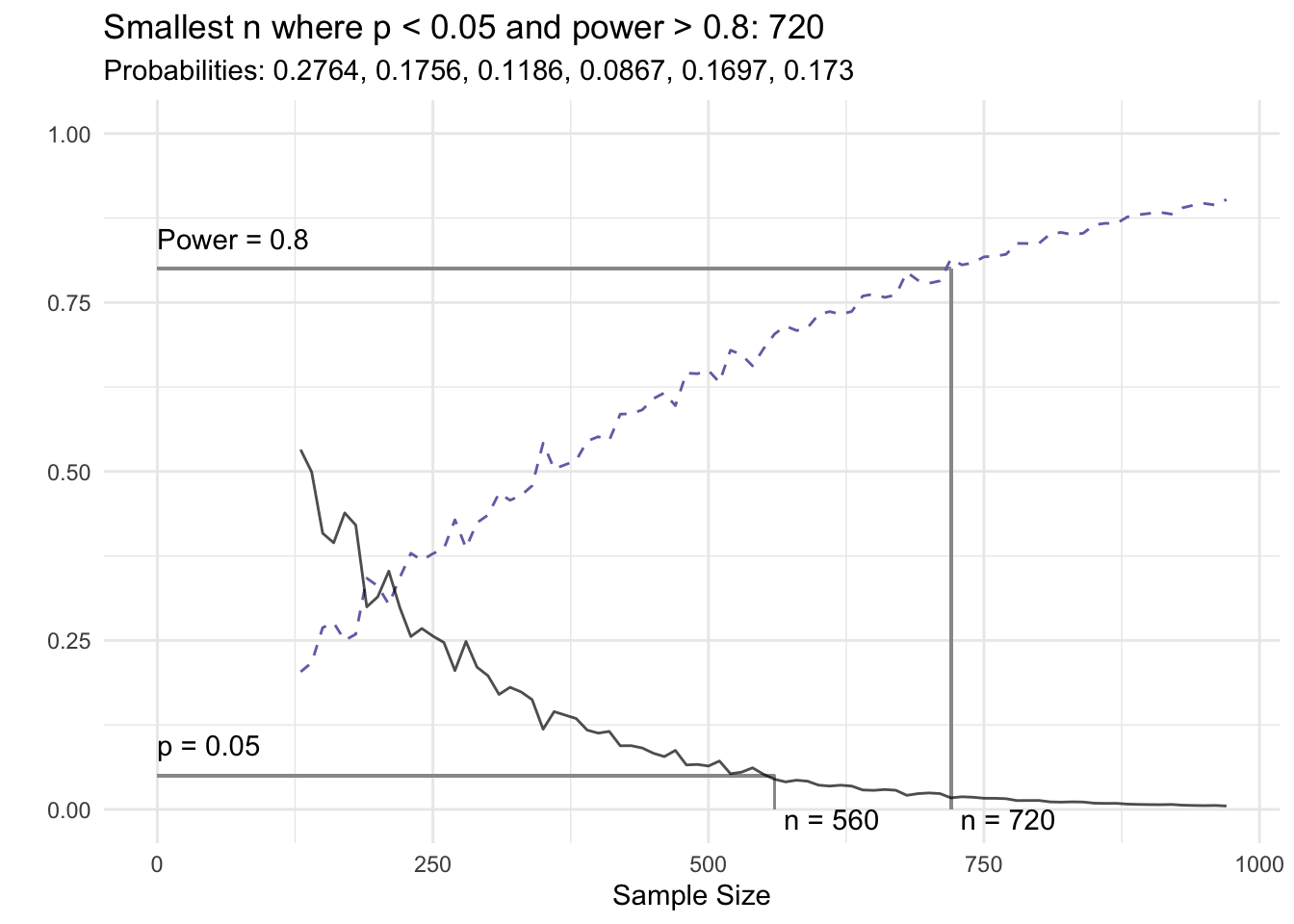
if(any(x$power > pow, na.rm = TRUE) & any(x$sig, na.rm = TRUE)) {
min_n <- min(x[x$sig & x$power > pow,]$n, na.rm = TRUE)
title <- paste0('Smallest n where p < ', attr(x, 'sig_level'), ' and power > ', pow, ': ',
prettyNum(min_n, big.mark = ','))
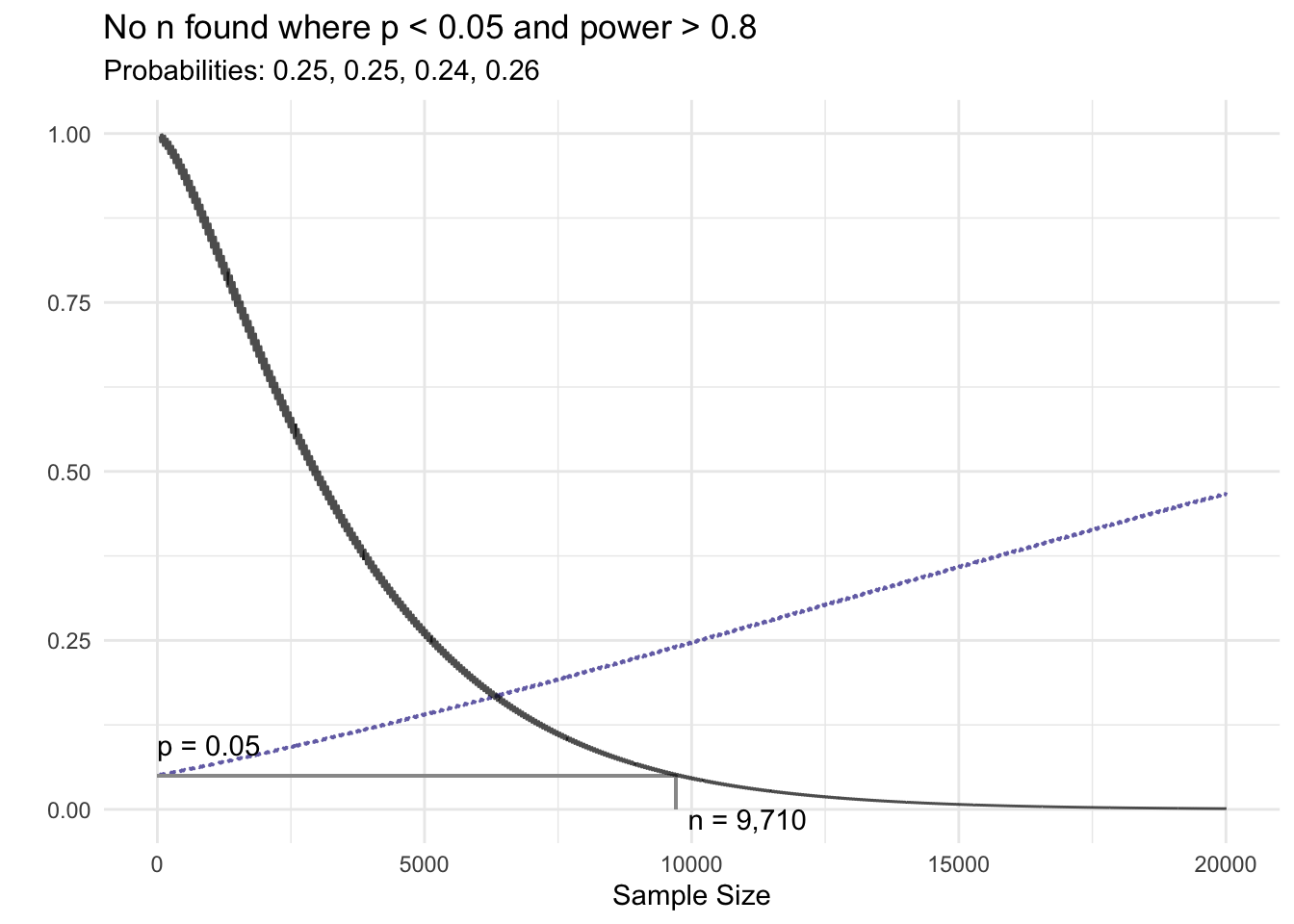
} else {
title <- paste0('No n found where p < ', attr(x, 'sig_level'), ' and power > ', pow)
}
}
p <- p +
ylim(c(0, 1)) +
ylab('') +
xlab('Sample Size') +
ggtitle(title,
subtitle = paste0('Probabilities: ', paste0(round(attr(x, 'probs'), digits = digits), collapse = ', ')))
return(p)
}